Can Frontier LLMs Predict Clinical Trial Outcomes?
Benchmarking Biological Alpha in Frontier LLMs.
tl;dr
Current Frontier LLMs are weak biological alpha generators.
LLM RCT-outcome prediction performance is driven by memorization, performance decreases with recency.
The field needs leakage-resistant bio-evals that genuinely test predictive validity on novel biology.
Chasing Predictive Validity
~90% of clinical programs fail. We still cannot reliably predict which biology will translate to humans. In practice, drug development remains a high-stakes capital allocation problem driven by expert judgment, imperfect experimental proxies, and uncertainty.
AI is promising to change that: from virtual cells, to virtual patients there is massive hype around models promising better biology. The reality is that, for most tools and evidence sources we don’t know how much valid information they add to the picture.
One of the tools promising change are the original heralds of the AI era: frontier LLMs. Frontier LLMs are increasingly validated on biology tasks, and biology has moved into the focus of frontier labs12. Already today, frontier LLMs are used in scientific workflows that affect real allocation decisions, including which therapeutic programs get advanced, partnered, or deprioritized.
This creates a practical and foundational question: can frontier LLMs truly assess the quality of biology being developed? Could they even assess translational performance of drug programs?
Established benchmarks such as TrialBench (2025) evaluate domain-specific architectures using static time splits. Here, we ask a different question: can off-the-shelf, general-purpose frontier models predict clinical trial outcomes in a way that plausibly reflects biological signal, rather than historical leakage?
The central challenge is train-test leakage through memorization. Real decisions happen at the frontier, where outcomes are not yet known. If performance is strongest on outcomes already disseminated in papers, registries, and media, then real prospective utility is limited.
We set up a straighforward experiment to test this.
Question: can frontier LLMs predict clinical trial outcomes, or are they mostly recovering already-known outcomes?
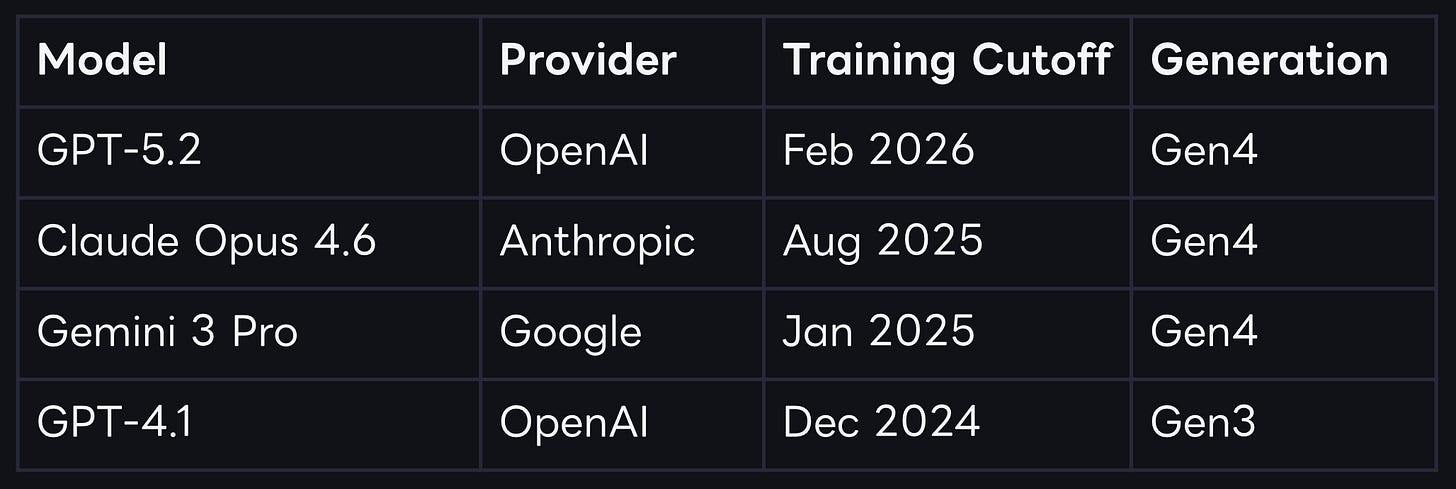
Experiment: test four frontier models (GPT-4.1, Gemini 3 Pro, Claude Opus 4.6, GPT-5.2) on clinical trial outcomes from ClinicalTrials.gov using a cutoff-centered evaluation design to clearly distinguish memorization from prospective prediction.
To the best of our knowledge, this specific assessment has not been done in a cutoff-aware way.
Setting up the experiment
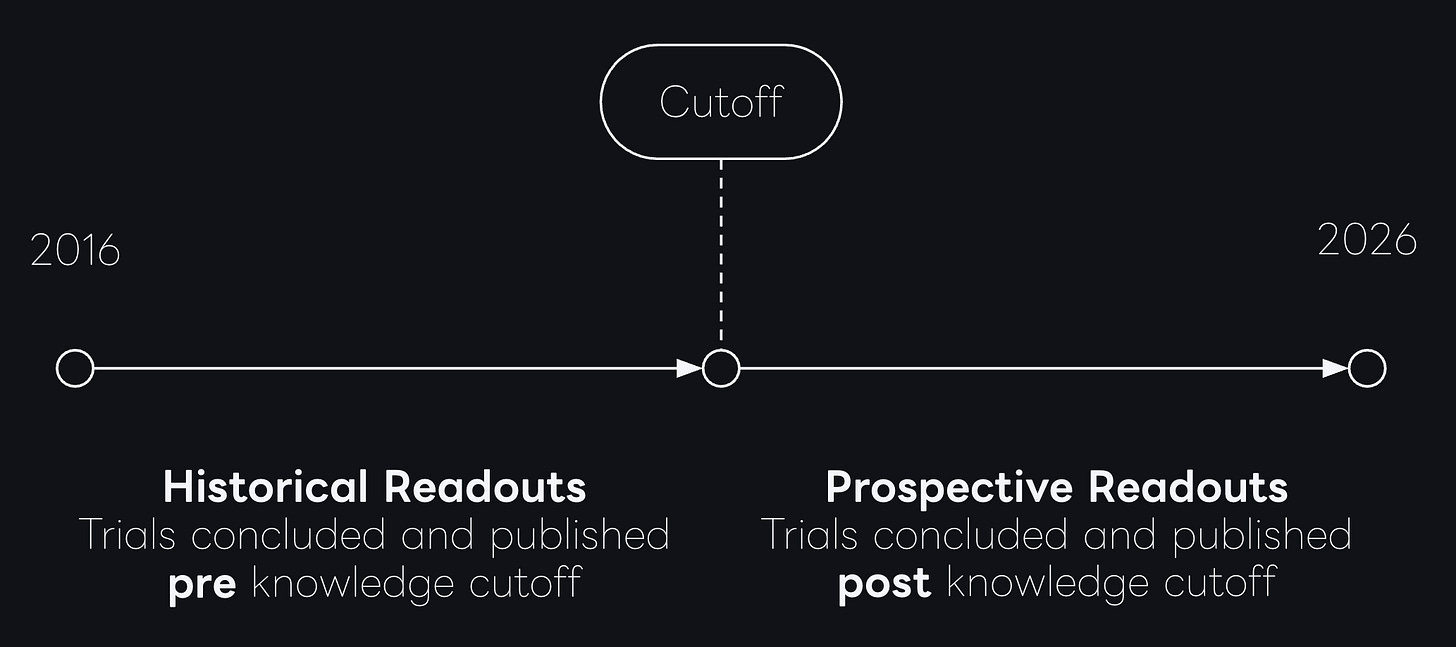
The core idea is pretty simple: Each model has a knowledge cutoff date, clinical trials that concluded before cutoff may have been represented in the model’s pretraining corpus, either directly or through derivative discussion. In high-visibility therapeutic areas, especially where mechanisms are repeatedly studied, historical outcomes can propagate broadly across scientific and non-scientific sources. Trials that concluded after cutoff should be less vulnerable to this effect and are the closest available proxy for true prospective prediction.
To set up the experiment, we first constructed a comprehensive time-stamped benchmark of clinical trials with known real world outcomes.
We ingested ClinicalTrials.gov, and extracted all randomized controlled trials between 2010-2016.
Next, we validated the trial completion status and the outcomes, leaving us with a set of 47,133 trials with validated clinical outcomes.
The specific outcome label was either
success (
primary endpoint met) orfailure
primary endpoint not met).
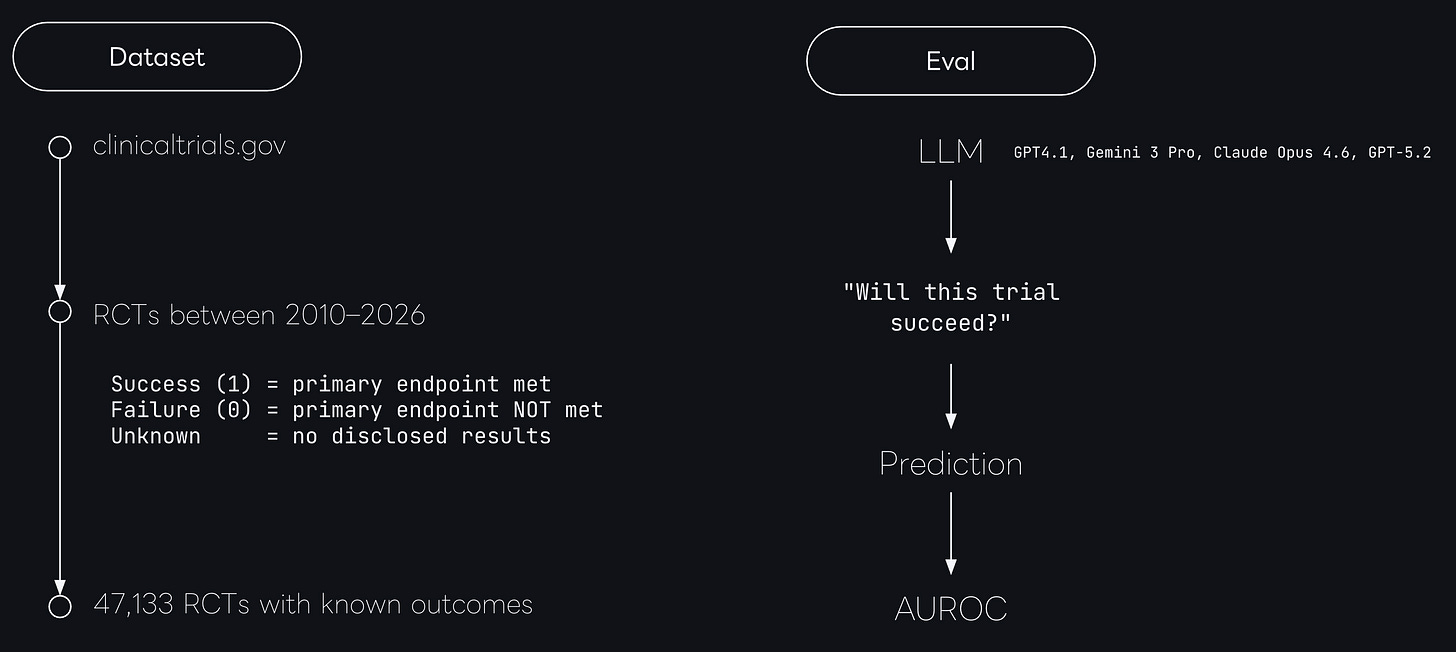
We then prompted each model to generate a probability of success score from 0 to 100 for all trials in our dataset3 and calculated the AUROC on the predictions.
Frontier models are good at picking winning mechanisms
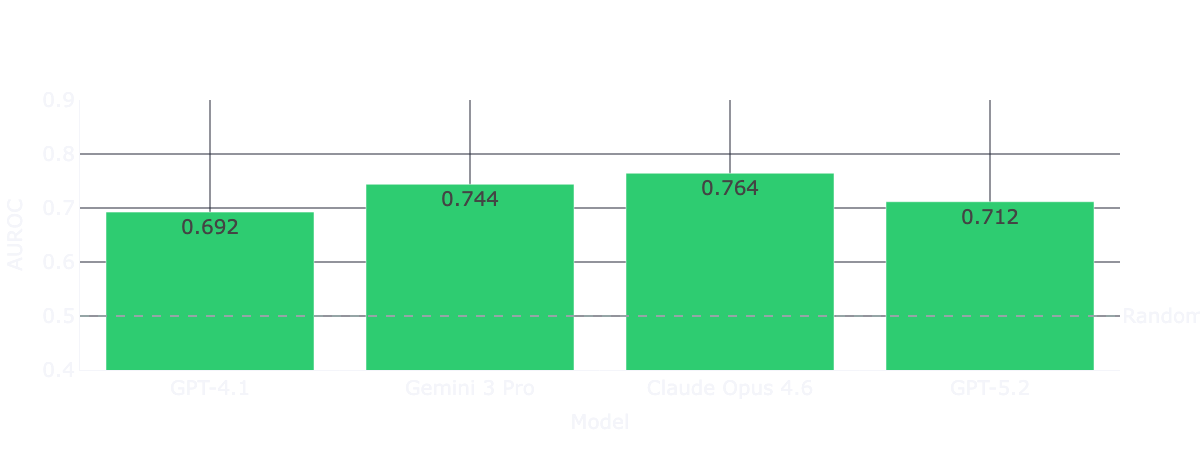
Across the full dataset of trials from 2016-2026, all four models deliver AUROC scores (0.69-0.76) on trial outcome classification.
At first, that looks pretty good. Recent benchmarks (such as TrialBench) reported AUROC scores in the 0.64–0.75 range for clinical trial approval predictions for domain-specific models under a conventional hold-out validation setting. This means naive frontier models are roughly matching heavily engineered, domain-specific architectures entirely out of the box.
But of course, this aggregate performance masks the source of signal. So next, we’re investigating historical vs prospective performance to control for train-test leakage.
Frontier models are good at memorizing winning mechanisms
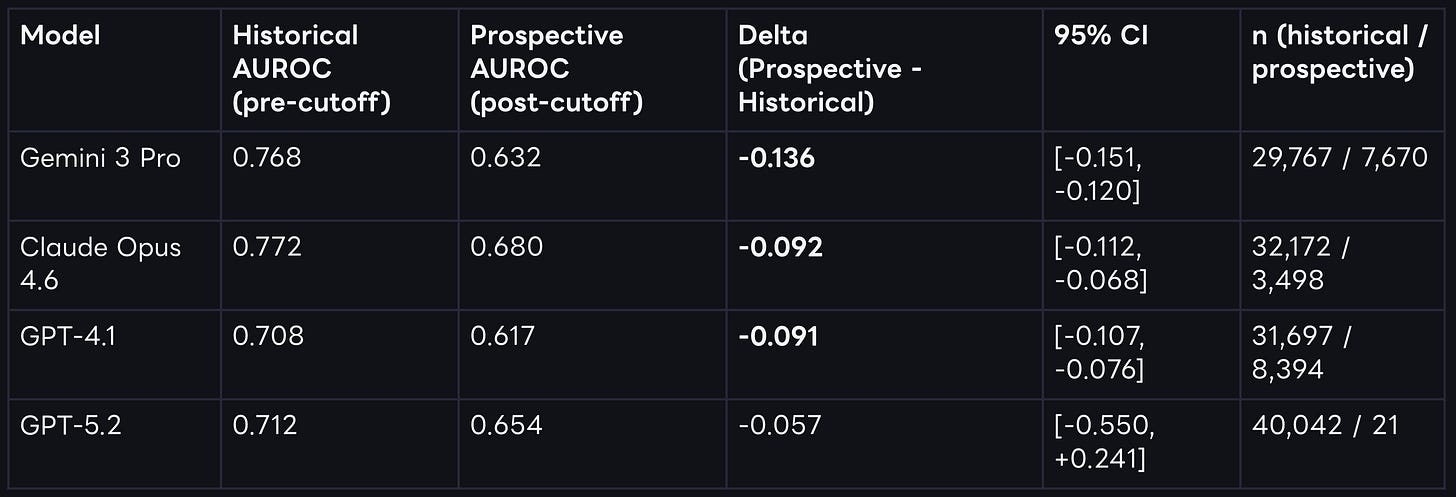
Controlling for train-test leakage through by computing two separate performance measures split by each model’s training cutoffs the picture looks entirely different. The temporally decomposed results show that performance drops in the prospective post-cutoff data for every single model tested.
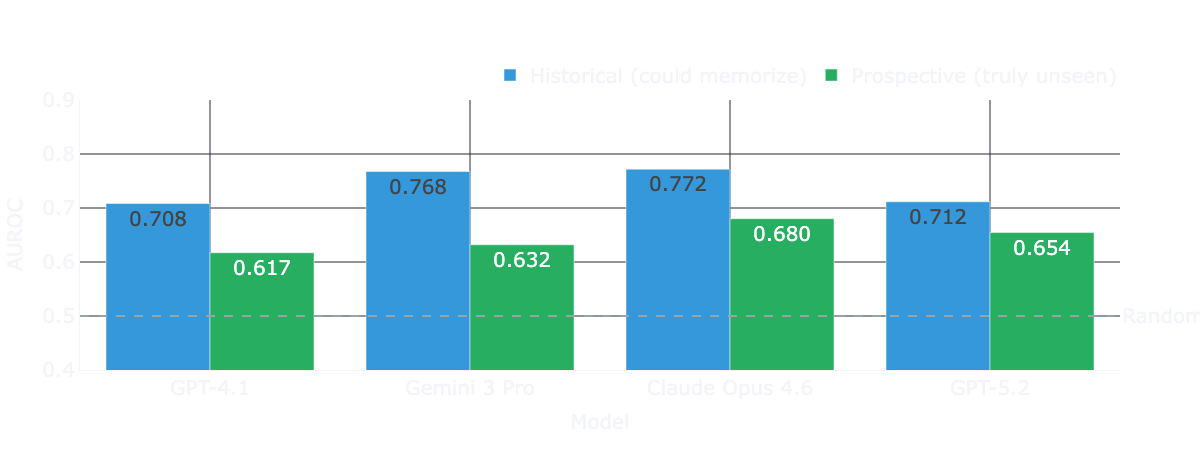
For three of the four models, this degradation is both large and statistically significant under bootstraped confidence intervals. We observe the largest drop for Gemini 3 Pro. For GPT-5.2, the directional drop is similar, but uncertainty remains high because only 21 prospective trials are available at this specific cutoff horizon.
While we see clear performance drops beyond cutoff dates for each model, the performance is not bad per se. Claude Opus 4.6 still shows an AUROC of 0.680 post cutoff - a pretty impressive result, without grounding, and a naive prompt.
However, drug programs take years to be developed, and each program leaves a decades-long paper trail before it reaches registrational trials. This paper trail of positive reporting, whether through published papers or data from previous phase trials, is almost certainly included in the models’ pre-training corpora.
We have no real way to test that directly. But one thing we can do is investigate if more recent trails, with potentially more recent biology, are assessed differently compared to data that has been around longer.
Performance weakens with recency
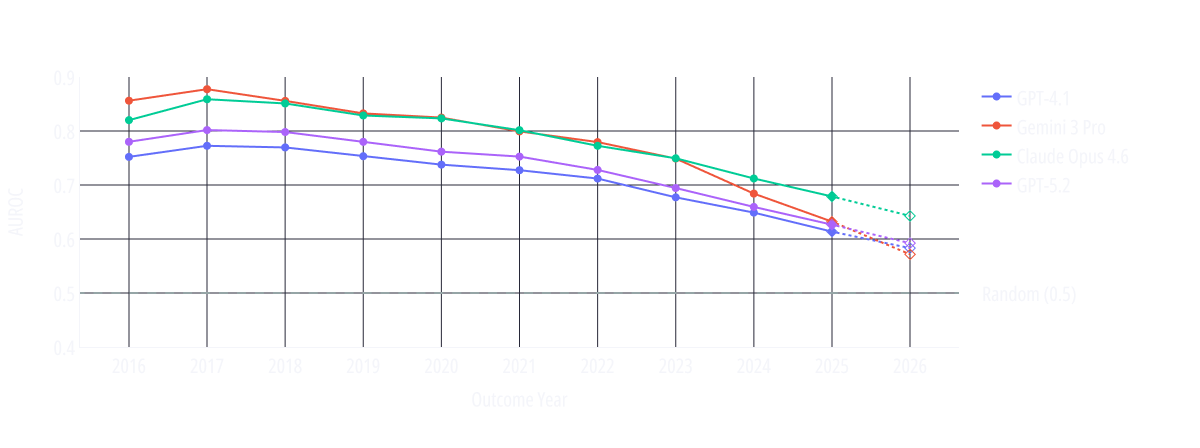
Rather than scoring models on a single pooled time axis, we aligned each model’s performance relative to its own cutoff date. We evaluated the AUROC per calendar year. This allows us to observe trends and run a fair comparison among models trained at different times.
The calendar-year resolved trend lines show AUROC declining steadily as the evaluation moves through time. Models generally perform weaker on more recent trials, supporting the memorization hypothesis.
If these systems possessed robust, mechanism-level predictive abilities independent of seen outcomes, their performance would be expected to remain comparatively stable across time. Instead, the performance decays.
What this decay means is that the task with greatest business relevance is the one with weakest evidence for high performance.
Trial-level examples: spike-testing memorization behavior
To illustrate this point, we decided to dive a bit deeper into the memorization angle and investigate some trial-level examples. Following the memorization paper-trail hypothesis, we expect pivotal trials with a large paper trail (i.e. global headlines, strong financial impact or guideline change), to result in predictions with extreme confidence.
The following cases were selected based on model confidence to avoid anecdotal cherry-picking:
NCT02021656: A 2015 trial investigating ledipasvir/sofosbuvir for HCV/HIV co-infection. This trial made global headlines
Outcome: Success (96% SVR12).
Model Performance: All four models assigned a probability of 96–98. GPT-4.1’s score of 98 sits at the 98.5th percentile of all its predictions, representing near-maximum confidence.
Market Context: The HCV direct-acting antiviral revolution was one of the most covered stories in modern medicine. Gilead, the sponsor, saw historic revenue growth and massive stock appreciation during this era as these curative treatments hit the market.
NCT04353037: This trial investigated hydroxychloroquine for COVID-19 and was terminated in 2021.
Outcome: Failure. It was terminated early after external data showed no clinical benefit, making global headlines.
Model Performance: All four models assigned a probability between 2 and 10. Gemini 3 Pro’s score of 2 sits at the 0.01st percentile, while GPT-4.1, GPT-5.2, and Opus 4.6 also ranked this in their bottom 1st percentile of predictions.
Context: The immense public and macroeconomic attention on COVID-19 therapeutics guaranteed this failure was heavily overrepresented in training data.
NCT02406027: This trial was terminated in 2018 due to safety concerns, specifically elevated liver enzymes and cognitive worsening.
Outcome: Failure.
Model Performance: Opus 4.6 assigned a probability of 5 (the 0.07th percentile). Gemini 3 Pro assigned 10, GPT-4.1 assigned 15, and GPT-5.2 assigned 20.
Market Context: Atabecestat, was a highly anticipated BACE inhibitor developed by Johnson & Johnson (Janssen) and Shionogi. The abrupt 2018 termination sent shockwaves through the sector and was a major catalyst in the subsequent collapse of the entire BACE inhibitor drug class.
While these examples do not prove memorization alone, they illustrate a broader pattern: high-confidence “predictions” on already-disseminated outcomes that received very high coverage in press and scientific journals.
What This Means
Before we move to interpretation, let’s summarize what we found:
Current frontier LLMs are strong retrievers of historical trial outcomes.
Out of the box performance on prospective trial readouts is much lower but still relevant (AUROC 0.680 for Claude Opus 4.6).
We have to assume that a substantial share of that capability is memorization from prior exposure to the paper trail leading up to the trial. Therefore, predictive reliability on truly novel programs remains unclear.
To be clear: this does not imply that LLMs have no role in drug development workflows.
It implies they should be used where recall-heavy capabilities are appropriate and memorization is not an issue. Many of these use cases exist in the current discovery workflows. However, that also mean that current frontierLLMs cannot (yet) be used as primary discovery engines where novelty and causal reasoning over unseen data are required.
Application in use cases, such as grading of novel biology, identification of novel targets or capital allocation decisions that assume model confidence equals out-of-sample validity, should be pursued with caution.
Practically, our findings demonstrate that naive LLMs are weak biological alpha generators. Because alpha depends entirely on information arbitrage not already embedded in the consensus, a system that primarily retrieves known outcomes is structurally limited as a predictive engine.
While our study successfully tests for the leakage of published trial results, testing for the predictive validity of truly novel biology faces an even more fundamental leakage problem. The standard scientific pipeline, where a paper is published, an LLM trains on that paper, a company initiates a related trial, and we evaluate the model on that trial, creating a chain that is virtually impossible to blind models against.
This raises an uncomfortable question for the field: is it even possible to design evaluations that genuinely assess a frontier LLM’s ability to predict truly novel biology?
We are actively working on this evaluation frontier. More to come, stay tuned!
https://www.anthropic.com/news/anthropic-partners-with-allen-institute-and-howard-hughes-medical-institute
https://openai.com/index/gpt-5-lowers-protein-synthesis-cost/
We iterated substantially on prompting and tested prompt variance; detailed prompt-ablation results will be shared another time.